By Ali Sundermier
Antibiotics have been a pillar of modern medicine since the 1940s. Streptomycin, which belongs to a class of antibiotics called aminoglycosides, was the first hint of light in the millennia-long search for a treatment for tuberculosis, which remains one of the deadliest infectious diseases in human history.
Today, aminoglycosides are the most commonly prescribed antibiotics in the world due to their low cost and high effectiveness in tackling a broad spectrum of bacterial infections. But they also bring along side effects that can have lifelong impacts. Depending on the dosage and the particular antibiotic, an estimated 10 to 20 percent of patients who take aminoglycosides suffer kidney damage and 20 to 60 percent end up with irreversible hearing loss.
Now researchers at the Department of Energy’s SLAC National Accelerator Laboratory have developed a new imaging technique to better understand the mechanisms that lead to hearing loss when aminoglycosides are introduced to the body. Using the lab’s Linac Coherent Light Source (LCLS) X-ray laser and Stanford Synchrotron Radiation Lightsource (SSRL), SLAC researchers, in collaboration with researchers at Stanford University, were able to observe interactions between the drugs and bacterial ribosomes at both extremely low and room temperatures, revealing never-before-seen details.
They also demonstrated how small modifications to the antibiotics can lead to dramatic changes in ribosome shape that eliminate hearing loss. The research could lead to a better understanding of which parts of a drug molecule cause unwanted reactions in the body, which will enable the development of more effective antibiotics with fewer side effects.
The group was led by research associate and senior author Hasan DeMirci. Their results were published in Nucleic Acids Research.
3D printing proteins
Hasan DeMirci refers to ribosomes – tiny molecular machines made up of tangles of RNA and proteins clumped together and intricately wired like ramen noodles in soup – as “the 3D printers of the human body.” The ribosomes synthesize proteins using the genetic information contained in DNA, “building our bodies from the ground up.”
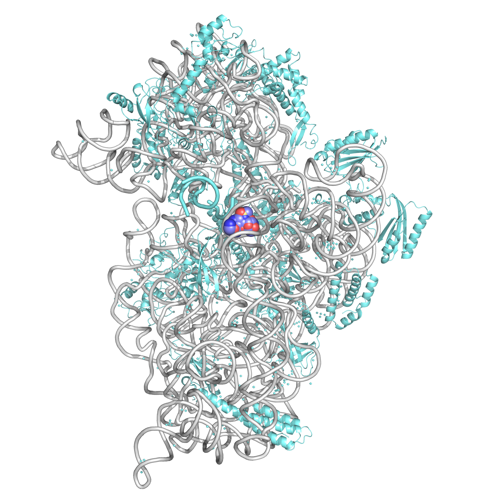
Ribosomes (shown here) are tiny molecular machines made up of tangles of RNA and proteins clumped together and intricately wired like ramen noodles in soup. (Hasan DeMirci/SLAC National Accelerator Laboratory)
“While one subunit of the ribosome, its brain, deciphers and translates the genetic code, the other, its hands, links together amino acids to form proteins,” DeMirci said.
Unlike viruses, which have to leech off hosts to survive, bacteria have their own ribosomes, which is where antibiotics come into play. Bacterial ribosomes are the targets of many antibiotics. So-called “cidal” antibiotics like aminoglycosides function by attacking the brains of bacterial ribosomes, causing them to make mistakes and fill the cells with protein-like garbage molecules.
“It's like a house with a lot of hoarded junk,” DeMirci says. “There's no going back. From that point the bacteria just die.”
The problem with this strategy is that human cells contain energy-producing factories called mitochondria that have their very own ribosomes – and since those ribosomes are dangerously similar to those found in bacteria, they’re also vulnerable to antibiotic attack.
“We're killing the bacteria, but the same drug gets into our mitochondria and destroys the ribosomes there,” DeMirci says. “Now we cannot produce those enzymes that power us. You take an antibiotic and you start losing your hearing, your kidney fails.”
Insights into molecular machinery
DeMirci has a strong interest in aminoglycosides because he can use them to gain insight into the molecular machinery of the ribosome.
“What I really want to know is what those drugs can teach us about how ribosomes decipher the genetic code,” DeMirci said. “Drugs give us an opportunity to stop that process at different stages to understand how each and every step is catalyzed by the ribosome.”
To better understand this process, he struck up a collaboration with Anthony Ricci, a biophysicist and professor of medicine at Stanford who focuses on the inner ear. In previous research, Ricci found that aminoglycosides infiltrate specialized channels to target the sensory cells essential to hearing.
“You can think of it as a roach motel,” Ricci says. “The drugs can get in but they can’t get out. They start to build up, binding to the ribosomes and altering protein synthesis. This puts a huge metabolic load on the sensory cells, which eventually leads to their deaths.”
A major goal of Ricci’s lab has been to design and develop new aminoglycosides that kill bacteria but cannot squeeze through the channel. In order to do this, the researchers need to understand exactly how the aminoglycosides interact with the ribosomes so they can modify parts of the drug without weakening its bacteria-killing properties.
Defrosting interactions
The best way to reach this understanding, researchers have found, is through a technique called X-ray crystallography. In X-ray crystallography, researchers use the patterns formed when a beam of X-rays scatters off a crystal sample to form a 3D model of how its atoms and molecules are arranged. This technique allows researchers to observe how a drug binds to a ribosome.
While the key interactions in these processes happen at body temperature, around 37 degrees Celsius, X-ray crystallography usually has to be done at extremely low, or cryogenic, temperatures, around minus 180 degrees Celsius. This leads to gaps in the data, obscuring tiny details that could greatly inform future experiments.
“Our bodies are warm, so the important biology is happening at body temperature,” DeMirci said, “but in crystallography everything is frozen. When you cool these processes down, you miss out on thermal fluctuations, tiny movements that could change your understanding of how the drugs and ribosomes are behaving.”
In order to design better antibiotics, they need to get as close a view as they can of this interaction happening under physiological conditions. At the LCLS, using a technique called serial femtosecond crystallography, DeMirci is able to catch the intricate waltz of the drugs and ribosomes at room temperature. Rather than freeze the ribosome crystals, the researchers suspend them in 'mother liquor,' a special chemical solution they were grown, so they are “swimming happily, still wiggling and fluctuating,” he says.
The crystals travel from a reservoir to the interaction region through a single capillary, like a garden hose. Once in the interaction region, the crystals are zapped with a beam of X-rays from the LCLS, which scatters off of them into a detector and provides the researchers with patterns they can use to build detailed 3D models of the ribosome before and after they’ve bound with the drugs. They then use these models to piece together a simulation of the interaction.
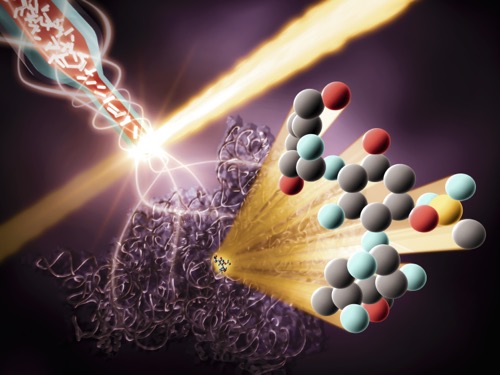
At LCLS, crystallized ribosomes travel through a capillary into the interaction region, where they are zapped with a beam of X-rays. The X-rays scatter off the crystals into a detector, providing the researchers with patterns they can use to build detailed 3D models of interactions between the drug and ribosome. (Greg Stewart/SLAC National Accelerator Laboratory).
Uncovering hidden wiggles
To demonstrate their technique, the researchers imaged modified and unmodified drugs binding to ribosomes at both cryogenic and room temperatures to see if they could catch any differences. They found that the drug molecules were less flexible at cryogenic temperatures: Tiny wiggles essential to a better understanding of their interactions with ribosomes were frozen in place.
“Despite the fact that we’ve recorded hundreds of thousands of structures of ribosomal interactions, less than a handful of new-generation drugs have been designed based on these cryogenic structures,” DeMirci said. “That’s because every small interaction makes a huge difference, even a single hydrogen bond.”
With the images taken at room temperature, Ricci’s group identified a site where the drug could be modified without altering its effectiveness.
“We now have some idea that when the drug binds with the ribosome, a global change occurs in the ribosome that might actually be important for the function of the antibiotic and the sensitivity of the ribosome,” Ricci said.
Refining the jigsaw pieces
In the next phase of experiments, DeMirci hopes to design a setup in which the antibiotics aren’t introduced until the last second before the ribosome is imaged so that they can watch as it binds to the ribosome, rather than just taking images before and after.
Up to this point, Ricci said, his group had been doing drug synthesis with very little information or insight into how the antibiotic interacts with the ribosome.
“What this paper and overall collaboration allow is a direct investigation of the drug-ribosome interaction,” he said. “It's like having more defined pieces to the jigsaw puzzle. You don't have to guess about what's happening.”
Developing antibiotics that can fight off drug-resistant bacteria with minimal side effects is essential because the rise of antibiotic resistant strains is currently the biggest threat to modern medicine, DeMirci said.
“Every year more than a million people die from tuberculosis and nearly half a million are HIV positive,” he said. “People don’t usually die from HIV or cancer, they die because their immune system is suppressed and they can’t fight off bacterial infections. That’s when you need antibiotics. But what if you don't have one that's effective against the resistant strains? That's exactly what's happening right now. This research can help us make informed decisions when designing the next generation of drugs.”
The research team included scientists from LCLS; SSRL; SLAC’s Biosciences Division; the Stanford PULSE Institute; and the Stanford School of Medicine.
LCLS and SSRL are DOE Office of Science user facilities. This work was supported by the National Institutes of Health, National Science Foundation, and Stanford.
Citation: O’Sullivan, Poitevin, Sierra, Gati et al., Nucleic Acids Research, 3 August 2018 (10.1093/nar/gky693)
Credit: SLAC National Accelerator Laboratory.
SLAC is a multi-program laboratory exploring frontier questions in photon science, astrophysics, particle physics and accelerator research. Located in Menlo Park, Calif., SLAC is operated by Stanford University for the U.S. Department of Energy's Office of Science.
SLAC National Accelerator Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time.
